Gene conversion
Gene conversion is the process by which one DNA sequence replaces a homologous sequence such that the sequences become identical after the conversion.[1] Gene conversion can be either allelic, meaning that one allele of the same gene replaces another allele, or ectopic, meaning that one paralogous DNA sequence converts another.[2]
Allelic gene conversion
[edit]Allelic gene conversion occurs during meiosis when homologous recombination between heterozygotic sites results in a mismatch in base pairing. This mismatch is then recognized and corrected by the cellular machinery causing one of the alleles to be converted to the other. This can cause non-Mendelian segregation of alleles in germ cells.[3]
Nonallelic/ectopic gene conversion
[edit]Recombination occurs not only during meiosis, but also as a mechanism for repair of double-strand breaks (DSBs) caused by DNA damage. These DSBs are usually repaired using the sister chromatid of the broken duplex and not the homologous chromosome, so they would not result in allelic conversion. Recombination also occurs between homologous sequences present at different genomic loci (paralogous sequences) which have resulted from previous gene duplications. Gene conversion occurring between paralogous sequences (ectopic gene conversion) is conjectured to be responsible for concerted evolution of gene families.[3][4][5]
Mechanism
[edit]
Conversion of one allele to the other is often due to base mismatch repair during homologous recombination: if one of the four chromatids during meiosis pairs up with another chromatid, as can occur because of sequence homology, DNA strand transfer can occur followed by mismatch repair. This can alter the sequence of one of the chromosomes, so that it is identical to the other.
Meiotic recombination is initiated through formation of a double-strand break (DSB). The 5’ ends of the break are then degraded, leaving long 3’ overhangs of several hundred nucleotides. One of these 3’ single stranded DNA segments then invades a homologous sequence on the homologous chromosome, forming an intermediate which can be repaired through different pathways resulting either in crossovers (CO) or noncrossovers (NCO). At various steps of the recombination process, heteroduplex DNA (double-stranded DNA consisting of single strands from each of the two homologous chromosomes which may or may not be perfectly complementary) is formed. When mismatches occur in heteroduplex DNA, the sequence of one strand will be repaired to bind the other strand with perfect complementarity, leading to the conversion of one sequence to another. This repair process can follow either of two alternative pathways as illustrated in the Figure. By one pathway, a structure called a double Holliday junction (DHJ) is formed, leading to the exchange of DNA strands. By the other pathway, referred to as Synthesis Dependent Strand Annealing (SDSA), there is information exchange but not physical exchange. Gene conversion will occur during SDSA if the two DNA molecules are heterozygous at the site of the recombinational repair. Gene conversion may also occur during recombinational repair involving a DHJ, and this gene conversion may be associated with physical recombination of the DNA duplexes on the two sides of the DHJ.
Biased vs. unbiased gene conversion
[edit]Biased gene conversion (BGC) occurs when one allele has a higher probability of being the donor than the other in a gene conversion event. For example, when a T:G mismatch occurs, it would be more or less likely to be corrected to a C:G pair than a T:A pair. This gives that allele a higher probability of transmission to the next generation. Unbiased gene conversion means that both possibilities occur with equal probability.
GC-biased gene conversion
[edit]GC-biased gene conversion (gBGC) is the process by which the GC content of DNA increases due to gene conversion during recombination.[4] Evidence for gBGC exists for yeasts and humans and the theory has more recently been tested in other eukaryotic lineages.[6] In analyzed human DNA sequences, crossover rate has been found to correlate positively with GC-content.[4] The pseudoautosomal regions (PAR) of the X and Y chromosomes in humans, which are known to have high recombination rates also have high GC contents.[3] Certain mammalian genes undergoing concerted evolution (for example, ribosomal operons, tRNAs, and histone genes) are very GC-rich.[3] It has been shown that GC content is higher in paralogous human and mouse histone genes that are members of large subfamilies (presumably undergoing concerted evolution) than in paralogous histone genes with relatively unique sequences.[7] There is also evidence for GC bias in the mismatch repair process.[3] It is thought that this may be an adaptation to the high rate of methyl-cytosine deamination which can lead to C→T transitions.
BGC of the Fxy gene in Mus musculus
[edit]The Fxy or Mid1 gene in some mammals closely related to house mice (humans, rats, and other Mus species) is located in the sex-linked region of the X chromosome. However, in Mus musculus, it has recently translocated such that the 3’ end of the gene overlaps with the PAR region of the X-chromosome, which is known to be a recombination hotspot. This portion of the gene has experienced a dramatic increase in GC content and substitution rate at the 3rd codon position as well as in introns but the 5’ region of the gene, which is X-linked, has not. Because this effect is present only in the region of the gene experiencing increased recombination rate, it must be due to biased gene conversion and not selective pressure.[4]
Impact of GC-biased gene conversion on human genomic patterns
[edit]GC content varies widely in the human genome (40–80%), but there seem to be large sections of the genome where GC content is, on average, higher or lower than in other regions.[3] These regions, although not always showing clear boundaries, are known as isochores. One possible explanation for the presence of GC-rich isochores is that they evolved due to GC-biased gene conversion in regions with high levels of recombination.
Evolutionary importance
[edit]Adaptive function of recombination
[edit]Studies of gene conversion have contributed to our understanding of the adaptive function of meiotic recombination. The ordinary segregation pattern of an allele pair (Aa) among the 4 products of meiosis is 2A:2a. Detection of infrequent gene conversion events (e.g. 3:1 or 1:3 segregation patterns during individual meioses) provides insight into the alternate pathways of recombination leading either to crossover or non-crossover chromosomes. Gene conversion events are thought to arise where the “A” and “a” alleles happen to be near the exact location of a molecular recombination event. Thus, it is possible to measure the frequency with which gene conversion events are associated with crossover or non-crossover of chromosomal regions adjacent to, but outside, the immediate conversion event. Numerous studies of gene conversion in various fungi (which are especially suited for such studies) have been carried out, and the findings of these studies have been reviewed by Whitehouse.[8] It is clear from this review that most gene conversion events are not associated with outside marker exchange. Thus, most gene conversion events in the several different fungi studied are associated with non-crossover of outside markers. Non-crossover gene conversion events are mainly produced by Synthesis Dependent Strand Annealing (SDSA).[9] This process involves limited informational exchange, but not physical exchange of DNA, between the two participating homologous chromosomes at the site of the conversion event, and little genetic variation is produced. Thus, explanations for the adaptive function of meiotic recombination that focus exclusively on the adaptive benefit of producing new genetic variation or physical exchange seem inadequate to explain the majority of recombination events during meiosis. However, the majority of meiotic recombination events can be explained by the proposal that they are an adaptation for repair of damage in the DNA that is to be passed on to gametes.[10]
Of particular interest, from the point of view that recombination is an adaptation for DNA repair, are the studies in yeast showing that gene conversion in mitotic cells is increased by UV[11][12] and ionizing radiation[13]
Genetic diseases of humans
[edit]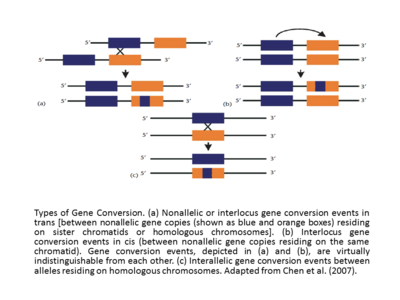
In the discussions of genetic diseases in humans, pseudogene mediated gene conversions that introduce pathogenic mutations into functional genes is a well known mechanism of mutation. In contrast, it is possible that pseudogenes could serve as templates. During the course of evolution, functional source genes which are potentially advantageous have been derived from multiple copies in their single source gene. The pseudogene-templated changes might eventually become fixed as long as they did not possess deleterious effects.[14] So, in fact, pseudogenes can act as sources of sequence variants which can be transferred to functional genes in novel combinations and can be acted upon by selection. Lectin 11 (SIGLEC11), a human immunoglobulin that binds to sialic acid, can be considered an example of such a gene conversion event which has played a significant role in evolution. While comparing the homologous genes of human SIGLEC11 and its pseudogene in the chimpanzee, gorilla and orangutan, it appears that there was gene conversion of the sequence of 5’ upstream regions and the exons that encode the sialic acid recognition domain, approximately 2kbp from the closely flanking hSIGLECP16 pseudogene (Hayakawa et al., 2005). The three pieces of evidence concerning this event have together suggested this as an adaptive change which is very evolutionarily important in genus Homo. Those includes that only in human lineage this gene conversion happened, the brain cortex has acquired an important expression of SIGLEC11 specifically in human lineage and the exhibition of a change in substrate binding in human lineage when compared to that of its counterpart in chimpanzees. Of course the frequency of the contribution of this pseudogene-mediated gene conversion mechanism to functional and adaptive changes in evolution of human is still unknown and so far it has been scarcely explored.[15] In spite of that, the introduction of positively selective genetic changes by such mechanism can be put forward for consideration by the example of SIGLEC11. Sometimes due to interference of transposable elements in to some members of a gene family, it causes a variation among them and finally it may also cease the rate of gene conversion due to lack of sequence similarity which leads to divergent evolution.
Genomic analysis
[edit]From various genome analyses, it was concluded that the double-strand breaks (DSB) can be repaired via homologous recombination by at least two different but related pathways.[14] In case of major pathway, homologous sequences on both sides of the DSB will be employed which seems to be analogous to the conservative DSB repair model [16] that was originally proposed for meiotic recombination in yeast.[17] where as the minor pathway is restricted to only one side of the DSB as postulated by nonconservative one-sided invasion model.[18] However, in both cases the sequence of the recombination partners will be absolutely conserved. By virtue of their high degree of homology, the new gene copies that came into existence following the gene duplication naturally tend to either unequal crossover or unidirectional gene conversion events. In the latter process, there exists the acceptor and donor sequences and the acceptor sequence will be replaced by a sequence copied from the donor, while the sequence of the donor remains unchanged.[15]
The effective homology between the interacting sequences makes the gene conversion event successful. Additionally, the frequency of gene conversion is inversely proportional to the distance between the interacting sequences in cis,[19][14] and the rate of gene conversion is usually directly proportional to the length of uninterrupted sequence tract in the assumed converted region. It seems that conversion tracts accompanying crossover are longer (mean length = ~460 bp) than conversion tracts without crossover (mean length = 55–290 bp).[20] In the studies of human globulin genes, it has long been supported that the gene conversion event or branch migration events can either be promoted or inhibited by the specific motifs that exist in the vicinity of the DNA sequence (Papadakis and Patrinos, 1999).[14] Another basic classification of gene conversion events is the interlocus (also called nonallelic) and interallelic gene conversions. The cis or trans nonallelic or interlocus gene conversion events occur between nonallelic gene copies residing on sister chromatids or homologous chromosomes, and, in case of interallelic, the gene conversion events take place between alleles residing on homologous chromosomes (Adapted from Chen et al., (2007).[15][14] If the interlocus gene conversion events are compared, it will be frequently revealed that they exhibit biased directionality. Sometimes, such as in case of human globin genes (Papadakis and Patrinos, 1999),[14] the gene conversion direction correlates with the relative expression levels of the genes that participate in the event, with the gene expressed at higher level, called the ‘master’ gene, converting that with lower expression, called the ‘slave’ gene. Originally formulated in an evolutionary context, the ‘master/slave gene’ rule should be explained with caution. In fact, the increase in gene transcription exhibits not only the increase in likelihood of it to be used as a donor but also as an acceptor.[14][21]
Effect
[edit]Normally, an organism that has inherited different copies of a gene from each of its parents is called heterozygous. This is generically represented as genotype: Aa (i.e. one copy of variant (allele) 'A', and one copy of allele 'a'). When a heterozygote creates gametes by meiosis, the alleles normally duplicate and end up in a 2:2 ratio in the resulting 4 cells that are the direct products of meiosis. However, in gene conversion, a ratio other than the expected 2A:2a is observed, in which A and a are the two alleles. Examples are 3A:1a and 1A:3a. In other words, there can, for example, be three times as many A alleles as a alleles expressed in the daughter cells, as is the case in 3A:1a.
Medical relevance
[edit]Gene conversion resulting in mutation of the CYP21A2 gene is a common underlying genetic cause of congenital adrenal hyperplasia. Somatic gene conversion is one of the mechanisms that can result in familial retinoblastoma, a congenital cancer of the retina, and it is theorized that gene conversion may play a role in the development of Huntington's disease.
References
[edit]- ^ Hastings, P.J. (2010-11-29). "Mechanisms of Ectopic Gene Conversion". Genes. 1 (3): 427–439. doi:10.3390/genes1030427. ISSN 2073-4425. PMC 3758752. PMID 24000309.
- ^ Trombetta, Beniamino; Fantini, Gloria; D’Atanasio, Eugenia; Sellitto, Daniele; Cruciani, Fulvio (2016-06-27). "Evidence of extensive non-allelic gene conversion among LTR elements in the human genome". Scientific Reports. 6 (1): 28710. Bibcode:2016NatSR...628710T. doi:10.1038/srep28710. ISSN 2045-2322. PMC 4921805. PMID 27346230.
- ^ a b c d e f Galtier N, Piganeau G, Mouchiroud D, Duret L (October 2001). "GC-content evolution in mammalian genomes: the biased gene conversion hypothesis". Genetics. 159 (2): 907–911. doi:10.1093/genetics/159.2.907. PMC 1461818. PMID 11693127.
- ^ a b c d Duret L, Galtier N (2009). "Biased gene conversion and the evolution of mammalian genomic landscapes". Annual Review of Genomics and Human Genetics. 10: 285–311. doi:10.1146/annurev-genom-082908-150001. PMID 19630562.
- ^ Harpak A, Lan X, Gao Z, Pritchard JK (November 2017). "Frequent nonallelic gene conversion on the human lineage and its effect on the divergence of gene duplicates". Proceedings of the National Academy of Sciences of the United States of America. 114 (48): 12779–12784. Bibcode:2017PNAS..11412779H. doi:10.1073/pnas.1708151114. PMC 5715747. PMID 29138319.
- ^ Pessia E, Popa A, Mousset S, Rezvoy C, Duret L, Marais GA (2012). "Evidence for widespread GC-biased gene conversion in eukaryotes". Genome Biology and Evolution. 4 (7): 675–682. doi:10.1093/gbe/evs052. PMC 5635611. PMID 22628461.
- ^ Galtier N (February 2003). "Gene conversion drives GC content evolution in mammalian histones". Trends in Genetics. 19 (2): 65–68. doi:10.1016/s0168-9525(02)00002-1. PMID 12547511.
- ^ Whitehouse HL (1982). Genetic Recombination: understanding the mechanisms. Wiley. p. 321 & Table 38. ISBN 978-0471102052.
- ^ McMahill MS, Sham CW, Bishop DK (November 2007). "Synthesis-dependent strand annealing in meiosis". PLOS Biology. 5 (11): e299. doi:10.1371/journal.pbio.0050299. PMC 2062477. PMID 17988174.
- ^ Bernstein H, Bernstein C, Michod RE (2011). "19. Meiosis as an Evolutionary Adaptation for DNA Repair". In Kruman I (ed.). DNA Repair. InTech. doi:10.5772/25117. ISBN 978-953-307-697-3. S2CID 32156088.
- ^ Ito T, Kobayashi K (October 1975). "Studies on the induction of mitotic gene conversion by ultraviolet irradiation. II. Action spectra". Mutation Research. 30 (1): 43–54. Bibcode:1975MRFMM..30...43I. doi:10.1016/0027-5107(75)90251-1. PMID 1101053.
- ^ Hannan MA, Calkins J, Lasswell WL (1980). "Recombinagenic and mutagenic effects of sunlamp (UV-B) irradiation in Saccharomyces cerevisiae". Molecular & General Genetics. 177 (4): 577–580. doi:10.1007/bf00272666. PMID 6991864. S2CID 31023471.
- ^ Raju MR, Gnanapurani M, Stackler B, Martins BI, Madhvanath U, Howard J, et al. (September 1971). "Induction of heteroallelic reversions and lethality in Saccharomyces cerevisiae exposed to radiations of various LET ( 60 Co rays, heavy ions and - mesons) in air and nitrogen atmospheres". Radiation Research. 47 (3): 635–643. Bibcode:1971RadR...47..635R. doi:10.2307/3573356. JSTOR 3573356. PMID 5119583.
- ^ a b c d e f g Chen JM (2001). Gene Conversion in Evolution and Disease. Wiley. ISBN 9780470015902.
- ^ a b c Chen JM, Cooper DN, Chuzhanova N, Férec C, Patrinos GP (October 2007). "Gene conversion: mechanisms, evolution and human disease". Nature Reviews. Genetics. 8 (10): 762–775. doi:10.1038/nrg2193. PMID 17846636. S2CID 205484180.
- ^ Szostak JW, Orr-Weaver TL, Rothstein RJ, Stahl FW (May 1983). "The double-strand-break repair model for recombination". Cell. 33 (1): 25–35. doi:10.1016/0092-8674(83)90331-8. PMID 6380756. S2CID 39590123.
- ^ Ota T, Nei M (January 1995). "Evolution of immunoglobulin VH pseudogenes in chickens". Molecular Biology and Evolution. 12 (1): 94–102. doi:10.1093/oxfordjournals.molbev.a040194. PMID 7877500.
- ^ Belmaaza A, Chartrand P (May 1994). "One-sided invasion events in homologous recombination at double-strand breaks". Mutation Research. 314 (3): 199–208. doi:10.1016/0921-8777(94)90065-5. PMID 7513053.
- ^ Schildkraut E, Miller CA, Nickoloff JA (2005). "Gene conversion and deletion frequencies during double-strand break repair in human cells are controlled by the distance between direct repeats". Nucleic Acids Research. 33 (5): 1574–1580. doi:10.1093/nar/gki295. PMC 1065255. PMID 15767282.
- ^ Jeffreys AJ, May CA (February 2004). "Intense and highly localized gene conversion activity in human meiotic crossover hot spots". Nature Genetics. 36 (2): 151–156. doi:10.1038/ng1287. PMID 14704667.
- ^ Schildkraut E, Miller CA, Nickoloff JA (April 2006). "Transcription of a donor enhances its use during double-strand break-induced gene conversion in human cells". Molecular and Cellular Biology. 26 (8): 3098–3105. doi:10.1128/MCB.26.8.3098-3105.2006. PMC 1446947. PMID 16581784.
External links
[edit]- Gene+conversion at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- images: http://www.web-books.com/MoBio/Free/Ch8D4.htm Archived 2022-03-20 at the Wayback Machine and http://www.web-books.com/MoBio/Free/Ch8D2.htm Archived 2022-01-27 at the Wayback Machine
